Research
At the Yan Lab in the Yale School of Medicine's Department of Pathology, I studied the function of Endogenous Retroviruses (ERVs), ancient viral sequences often considered "junk DNA." My project focused on reactivating these ERVs through precise epigenetic editing to better understand their role in cancer and immunity.
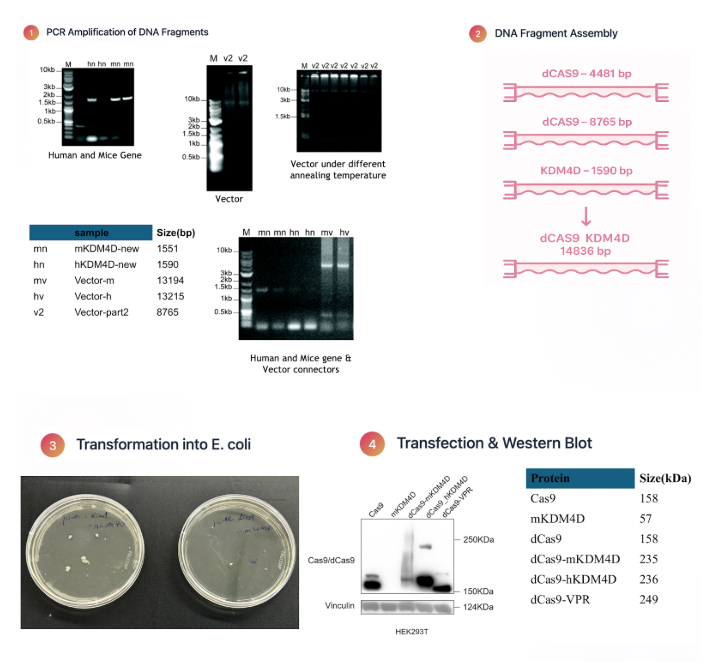
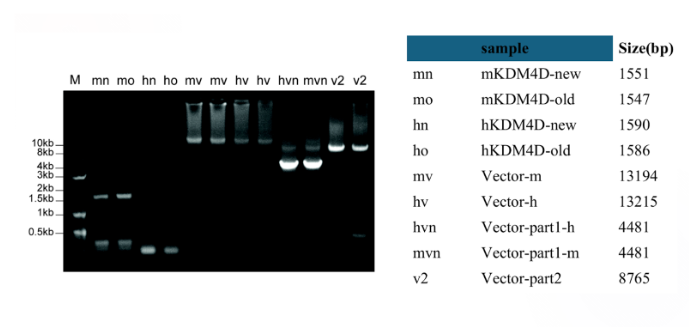
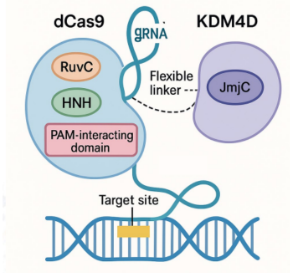
I constructed the lab's central molecular tool, a Cas9 KDM4D fusion protein designed to remove repressive H3K9me3 histone marks from ERV loci. I designed primers, performed PCR-based molecular cloning, and successfully assembled the lentiviral plasmid carrying our editor. After delivering the construct into target cells, I validated its expression through Western blot analysis.
This validated construct became the foundation for the lab's large-scale study on ERV function, enabling new exploration into how ERV reactivation may influence tumor biology and immune responses. I concluded my research by presenting my methods and results at a formal Yan Lab meeting at Yale.
Website of the lab: https://medicine.yale.edu/lab/yan/research/
Research: Melanoma and Endogenous Retroviruses
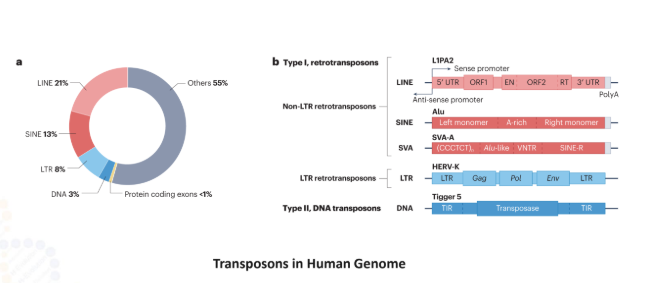
Endogenous Retroviruses (ERVs) are ancient viral elements that make up nearly 8% of the human genome. Once infectious, these sequences are now silenced by epigenetic modifications such as H3K9me3 to preserve genomic stability. Recent studies suggest that reactivation of ERVs in tumor cells can mimic viral infection and stimulate strong immune responses against cancer. This phenomenon, known as viral mimicry, represents an emerging direction in melanoma immunotherapy research.
At the Yale School of Medicine, the Yan Laboratory studies how histone demethylases regulate ERV silencing and how their dysregulation affects cancer progression and immune response. In this context, I contributed to the lab's ongoing effort to understand the functional role of ERV reactivation through precise epigenetic editing.
Project Focus: Locus-Specific Reactivation of ERVs
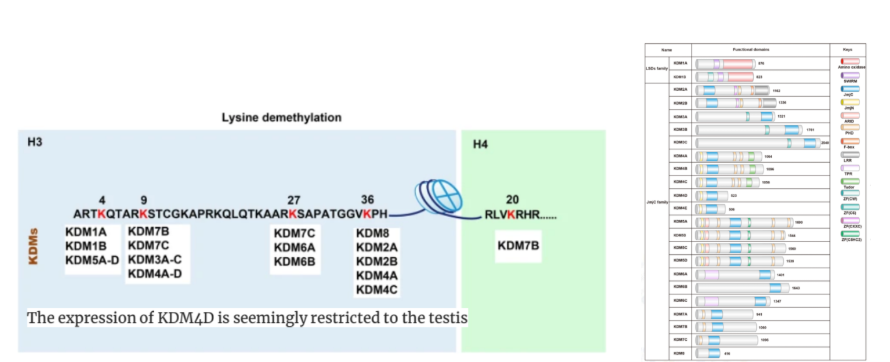
My research focused on developing a molecular tool to specifically reactivate ERVs by removing repressive histone marks. I constructed a dCas9-KDM4D fusion protein, engineered to erase H3K9me3 at ERV loci without altering the rest of the genome. To achieve this, I designed primers and performed PCR-based molecular cloning to assemble the lentiviral plasmid carrying the fusion construct.
After verifying correct assembly through colony screening and sequencing, I transfected mammalian cells and confirmed expression of the fusion protein by Western blot analysis. This validated system now serves as the foundation for the lab's functional screening platform that will map which ERV families most strongly influence anti-tumor immunity in melanoma.
Research Impact
The successful construction and validation of the dCas9-KDM4D epigenetic editor enable the lab to study ERV activation with unprecedented precision. This tool provides a framework for exploring how epigenetic regulation of viral elements shapes the tumor microenvironment and immune signaling.
By contributing to this project, I helped establish a critical molecular platform that supports the lab's broader goal of uncovering how chromatin modifications influence cancer immunity. The insights gained may ultimately inform new therapeutic strategies that harness the body's immune system to recognize and eliminate tumor cells.